This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is Congenital Nystagmus?
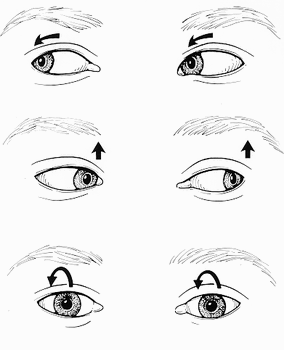
Figure 1. Eye movement of nystagmus
Congenital nystagmus is a condition where the afflicted person experiences rapid, uncontrollable eye movements. According to Medline Plus, these movements may be side to side (horizontal), up and down (vertical), or rotary (torsional). There are also two different classes of nystagmus, sensory nystagmus which is related to vision loss, and motor nystagmus which is related to the control of muscle function [1]. Occasionally, this condition can also develop later in life due to an accident or illness, but this form is not related to genetics. Infantile nystagmus is the most common form with an estimated prevalence of 1.9 cases in 1000 people [2]. Normally, affected people don’t notice the eye movements, especially if they are small. If they are larger however, nystagmus can cause the sharpness of vision to be less than 20/20 [Medline Plus]. What really makes infantile nystagmus unique is that it is not correlated with any other condition that may cause irregular eye movement.
The genetics of this disorder are tricky. A grand total of five genes have been identified to cause infantile nystagmus, three of which are autosomal, and two are X-linked [3]. Also, there have been cases of both dominant and recessive inheritance of nystagmus among different families. These conditions include ocular albinism, congenital stationary night blindness, blue cone monochromatism and sensory defects of early childhood [4]. Symptoms of nystagmus usually start between four and six months of age, and the condition is present in a diverse set of populations.
The genetics of this disorder are tricky. A grand total of five genes have been identified to cause infantile nystagmus, three of which are autosomal, and two are X-linked [3]. Also, there have been cases of both dominant and recessive inheritance of nystagmus among different families. These conditions include ocular albinism, congenital stationary night blindness, blue cone monochromatism and sensory defects of early childhood [4]. Symptoms of nystagmus usually start between four and six months of age, and the condition is present in a diverse set of populations.
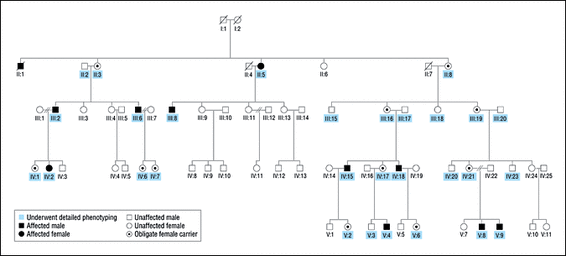
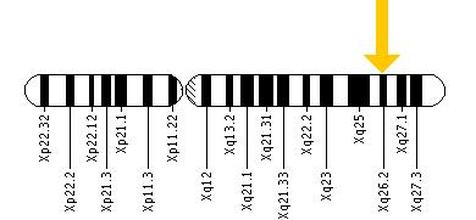
The gene most commonly studied for infantile nystagmus is the Ferm domain containing protein 7 (FRMD7) gene which is located on the X chromosome. The location of FRMD7 causes the severity of this condition to vary greatly, especially in women due to random X inactivation in the embryo. Since males only have one X chromosome, if the FRMD7 mutation exists on that chromosome, they will be affected. Females however, have two copies which cause a mosaic in their cells when one of those X’s is inactivated in the early embryo. This can cause less severe symptoms, and approximately half of the females with only one altered copy of the FRMD7 gene have no symptoms of nystagmus [3].
Treatment
There currently is no curative treatment for nystagmus, but there are a few techniques that help those with decreased sharpness of vision see more clearly. One of these techniques is finding he null point, or the gaze angle where nystagmus dampens. Often, this can be found by looking to one side so eye movement is reduced, and can even lead to abnormal posture in order to see more clearly [American Nystagmus Network]. There are also four surgeries which can sometimes help correct aspects of nystagmus, but those are often seen as a last resort [6]. The most common of these surgeries involves re-positioning of the nerves behind the eye. It is meant to trick the brain and slow down the nystagmus eye movement.
Below is a video describing one of the surgeries to help treat nystagmus symptoms.
Below is a video describing one of the surgeries to help treat nystagmus symptoms.
From YouTube 2012. Live Webcast: Treatment for Congenital Nystagmus. Retrieved from:
http://www.youtube.com/watch?v=ZOU7QlwsUJM&feature=relmfu
FRMD7
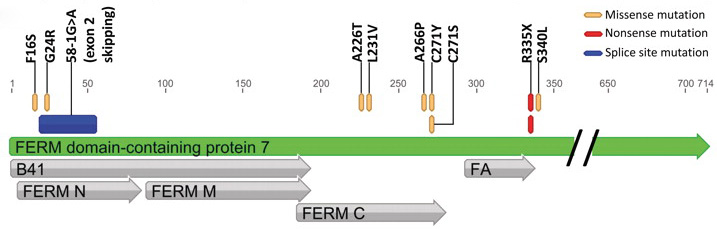
FRMD7 contains a FERM domain, which is named for the four original proteins in which it was originally described. This FERM domain contains three subdomains that interact to form a single molecule. This domain is usually related with proteins associated with the cytoskeleton. In particular, the FERM domain links actin filaments and adhesion proteins in the cytoskeleton [7].
FRMD7 itself has 12 exons which encode a 714 polypeptide chain [Accession Number: NP_919253]. The specific function of FRMD7 is not well understood but several studies suggest it plays an important role in neuronal development, particularly in regions associated with ocular movement. The most common mutations in FRMD7 are point missense mutations that cause an amino acid change at that location. Figure 4 also shows a mutation where exon 2 is skipped due to an introduced splice site on the primary mRNA transcript with a point mutation [3].
In both the human and mouse embryonic brains, FRMD7 is specially and temporally regulated, and in culture the protein is strongly associated with the growth cone of developing neurons [8]. What this means is, a mutation in the FRMD7 protein can lead to stunted axon growth and immature neuron cells. This can lead directly to infantile nystagmus since FRMD7 is associated with ocular movement.
References
[1] Windsor, R., Windsor, L. Understanding Nystagmus. Low Vision Centers of Indiana. Retrieved February 1, 2012, from http://www.lowvision.org/nystagmus.htm
[2] Watkins, R., Thomas, M. G., Talbot, C. J. Gottlob, I., Shakelton, S. (2012)
The role of FRMD7 in Idiopathic Infantile Nystagmus. Journal of Opthalmology, 2012(2012), 460956. doi: 10.1155/2012/460956
[3] Thomas, M. G., Crosier, M., Lindsay, S., Kumar, A., Thomas, S., Araki, M., Talbot, C., McLean, R. J., Surendran, M., Taylor, K., Leroy, B. P., Moore, A. T., Hunter, D. G., Hertle, R. W., Tarpey, P., Langmann, A., Lindner, S., Brandner, M., Gottlob., I. (2011)
The clinical and molecular genetic features of idiopathic infantile periodic alternating nystagmus. Brain, a Journal of Neurology, 2011(134), 892, doi: 10.1093/brain/awq373
[4] Tarpey P., Thomas S., Sarvananthan N., Mallya U., Lisgo S., Talbot C. J., Roberts E. O., Awan M., Surendran M., McLean R. J., Reinecke R. D., Langmann A., Lindner S., Koch M., Jain S., Woodruff G., Gale R. P., Degg C., Droutsas K., Asproudis I., Zubcov A. A., Pieh C., Veal C. D., Machado R. D., Backhouse O. C., Baumber L., Constantinescu C. S., Brodsky M. C., Hunter D. G., Hertle R. W., Read R. J., Edkins S., O'Meara S., Parker A., Stevens C., Teague J., Wooster R., Futreal P. A., Trembath R. C., Stratton M. R., Raymond F. L., Gottlob I. (2006)
Mutations in FRMD7, a newly identified member of the FERM family, cause X-linked idiopathic congenital nystagmus. Nature Genetics, 38, 1242. doi: 10.1038/ng1893
[5] Self, J. E., Shawkat, F., Malpas C. T., Thomas S., Harris, C. M., Hodgkings P. R., Chen, X., Trump, D., Lotery, A. J. (2007)
Allelic variation of the FRMD7 gene in congenital idiopathic nystagmus. Ophthalmic Molecular Genetics, 125(9), 1255. Retrieved from: http://archopht.ama-assn.org/cgi/content/full/125/9/1255
[6] Lee, J. (2002)
Surgical management of nystagmus. Journal of the Royal Society of Medicine, 95(5), 238. Retrieved from: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1279676/
[7] Hamada, K., Shimizu, T., Matsui, T., Tsukita, S., Tsukita, S., Hakoshima, T. (2000)
Structural basis of the membrane-targeting and unmasking mechanisms of the radixin FERM domain. European Molecular Biology Organization Journal, 19(2000), 4449. doi: 10.1093/emboj/19.17.4449
[8] Betts-Henderson J., Bartesaghi, S., Crosier, M., Lindsay, S., Chen, H., Salomoni, P., Gottlob, I., Nicotera P. (2010)
The nystagmus-associated FRMD7 gene regulates neuronal outgrowth and development. Human Molecular Genetics, 19(2), 342. doi: 10.1093/hmg/ddp500
[2] Watkins, R., Thomas, M. G., Talbot, C. J. Gottlob, I., Shakelton, S. (2012)
The role of FRMD7 in Idiopathic Infantile Nystagmus. Journal of Opthalmology, 2012(2012), 460956. doi: 10.1155/2012/460956
[3] Thomas, M. G., Crosier, M., Lindsay, S., Kumar, A., Thomas, S., Araki, M., Talbot, C., McLean, R. J., Surendran, M., Taylor, K., Leroy, B. P., Moore, A. T., Hunter, D. G., Hertle, R. W., Tarpey, P., Langmann, A., Lindner, S., Brandner, M., Gottlob., I. (2011)
The clinical and molecular genetic features of idiopathic infantile periodic alternating nystagmus. Brain, a Journal of Neurology, 2011(134), 892, doi: 10.1093/brain/awq373
[4] Tarpey P., Thomas S., Sarvananthan N., Mallya U., Lisgo S., Talbot C. J., Roberts E. O., Awan M., Surendran M., McLean R. J., Reinecke R. D., Langmann A., Lindner S., Koch M., Jain S., Woodruff G., Gale R. P., Degg C., Droutsas K., Asproudis I., Zubcov A. A., Pieh C., Veal C. D., Machado R. D., Backhouse O. C., Baumber L., Constantinescu C. S., Brodsky M. C., Hunter D. G., Hertle R. W., Read R. J., Edkins S., O'Meara S., Parker A., Stevens C., Teague J., Wooster R., Futreal P. A., Trembath R. C., Stratton M. R., Raymond F. L., Gottlob I. (2006)
Mutations in FRMD7, a newly identified member of the FERM family, cause X-linked idiopathic congenital nystagmus. Nature Genetics, 38, 1242. doi: 10.1038/ng1893
[5] Self, J. E., Shawkat, F., Malpas C. T., Thomas S., Harris, C. M., Hodgkings P. R., Chen, X., Trump, D., Lotery, A. J. (2007)
Allelic variation of the FRMD7 gene in congenital idiopathic nystagmus. Ophthalmic Molecular Genetics, 125(9), 1255. Retrieved from: http://archopht.ama-assn.org/cgi/content/full/125/9/1255
[6] Lee, J. (2002)
Surgical management of nystagmus. Journal of the Royal Society of Medicine, 95(5), 238. Retrieved from: http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1279676/
[7] Hamada, K., Shimizu, T., Matsui, T., Tsukita, S., Tsukita, S., Hakoshima, T. (2000)
Structural basis of the membrane-targeting and unmasking mechanisms of the radixin FERM domain. European Molecular Biology Organization Journal, 19(2000), 4449. doi: 10.1093/emboj/19.17.4449
[8] Betts-Henderson J., Bartesaghi, S., Crosier, M., Lindsay, S., Chen, H., Salomoni, P., Gottlob, I., Nicotera P. (2010)
The nystagmus-associated FRMD7 gene regulates neuronal outgrowth and development. Human Molecular Genetics, 19(2), 342. doi: 10.1093/hmg/ddp500
Site created by: Kristen Klimo
Last updated: 5/11/12
University of Wisconsin-Madison
Last updated: 5/11/12
University of Wisconsin-Madison