This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Model Organisms
In genetics, it's extremely common for researchers to use model organisms to conduct experiments when experimentation on humans is unethical or impossible. This use of model organisms is possible because of evolution. As organisms diverged over time, metabolic pathways, genetic material and and developmental pathways were conserved [1]. Model organisms are often chose for their short life cycle, ability to be manipulated and non-specialist living requirements. Also, researchers also tend to choose organisms whose genomes are already sequenced. Common model organisms in genetics are C. elegans, Drosophila (fruit flies) and mice.
When choosing which model organism, it is important to consider the trait being studies. For nystagmus, a model organism with eyes similar to human eyes would need to be chosen, in order to properly study defects in the protein and genome. This automatically eliminates C. elegans and Drosophila since C. elegans don't have any eyes, and Drosophila have a compound eye that is not a good model for humans. This leaves mice.
The mouse model organism is the most closely related to humans. The two genomes are approximately the same size which contain the same number of genes. They also show extensive synteny, meaning the same gene order. Additionally, mutations that cause diseases in humans often cause similar diseases in mice [2]. Mice are also useful because they have a short breeding cycle, and are capable of producing 10 or more offspring per litter. They are also very easy to mutate and maintain mutant inbred strains.
When choosing which model organism, it is important to consider the trait being studies. For nystagmus, a model organism with eyes similar to human eyes would need to be chosen, in order to properly study defects in the protein and genome. This automatically eliminates C. elegans and Drosophila since C. elegans don't have any eyes, and Drosophila have a compound eye that is not a good model for humans. This leaves mice.
The mouse model organism is the most closely related to humans. The two genomes are approximately the same size which contain the same number of genes. They also show extensive synteny, meaning the same gene order. Additionally, mutations that cause diseases in humans often cause similar diseases in mice [2]. Mice are also useful because they have a short breeding cycle, and are capable of producing 10 or more offspring per litter. They are also very easy to mutate and maintain mutant inbred strains.
Mice as a Model for Nystagmus
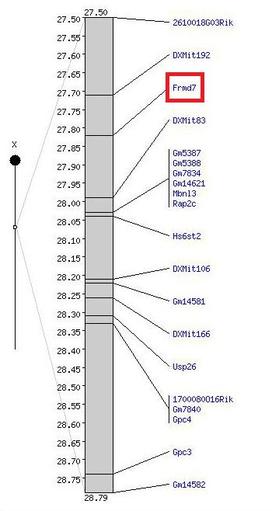
Figure 1. FRMD7 gene on mouse X chromosome
For this analysis, the database Mouse Genome Informatics (MGI) was used. According to their website, MGI provides integrated access to data on the genetics, genomics, and biology of the labratory mouse. This is helpful because it eliminates the need to search a more generalized search engine for accurate results. The figure to the left, shows the location of the FRMD7 gene on the mouse X chromosome, indicated by the red box.
The MGI analysis also showed that the gene ontology in mice also matched the human gene. The mouse FRMD7 is involved in growth cone projection, neuronal development and cytoskeleton activities, among other things. Since these functions are identical to human FRMD7, this reinforces the mouse as a very useful model organism for studying nystagmus.
The MGI analysis also showed that the gene ontology in mice also matched the human gene. The mouse FRMD7 is involved in growth cone projection, neuronal development and cytoskeleton activities, among other things. Since these functions are identical to human FRMD7, this reinforces the mouse as a very useful model organism for studying nystagmus.
Analysis and Discussion
To date there is no information about the complete FRMD7 gene or protein in either WormBase for C. elegans, or FlyBase for Drosophila. However, when the protein domains were searched separately, FERM C, yielded results in FlyBase. FERM C is the location most commonly associated with nystagmus-causing mutations in the gene [3].
One thing to note is the presense of the FRMD7 protein in C. elegans. It may be worth exploring what this gene is doing in that model system in order to create RNAi experiments in C. elegans, since FRMD7 is still so well conserved in an organism that does not have eyes. For more information on this topic, please visit the Conclusions and Future Directions page of this website.
References
[1] Fox, M. A. (1986)
The Case for Animal Experimentation: An Evolutionary and Ethical Perspective. Retrieved May 21, 2012, from http://books.google.com/books?id=yTfNH3cScKAC&printsec=frontcover&source=gbs_ge_summary_r&cad=0#v=onepage&q&f=false
[2] Twyman, R. (2002)Model Organisms: The Mouse. Retrieved May 21, 2012 from http://genome.wellcome.ac.uk/doc_WTD020804.html
[3] Thomas, M. G., Crosier, M., Lindsay, S., Kumar, A., Thomas, S., Araki, M., Talbot, C., McLean, R. J., Surendran, M., Taylor, K., Leroy, B. P., Moore, A. T., Hunter, D. G., Hertle, R. W., Tarpey, P., Langmann, A., Lindner, S., Brandner, M., Gottlob., I. (2011)
The clinical and molecular genetic features of idiopathic infantile periodic alternating nystagmus. Brain, a Journal of Neurology, 2011(134), 892, doi: 10.1093/brain/awq373
The Case for Animal Experimentation: An Evolutionary and Ethical Perspective. Retrieved May 21, 2012, from http://books.google.com/books?id=yTfNH3cScKAC&printsec=frontcover&source=gbs_ge_summary_r&cad=0#v=onepage&q&f=false
[2] Twyman, R. (2002)Model Organisms: The Mouse. Retrieved May 21, 2012 from http://genome.wellcome.ac.uk/doc_WTD020804.html
[3] Thomas, M. G., Crosier, M., Lindsay, S., Kumar, A., Thomas, S., Araki, M., Talbot, C., McLean, R. J., Surendran, M., Taylor, K., Leroy, B. P., Moore, A. T., Hunter, D. G., Hertle, R. W., Tarpey, P., Langmann, A., Lindner, S., Brandner, M., Gottlob., I. (2011)
The clinical and molecular genetic features of idiopathic infantile periodic alternating nystagmus. Brain, a Journal of Neurology, 2011(134), 892, doi: 10.1093/brain/awq373
Site created by: Kristen Klimo
Last updated: 3/22/2012
University of Wisconsin-Madison
Last updated: 3/22/2012
University of Wisconsin-Madison
