This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Protein-Protein Interactions
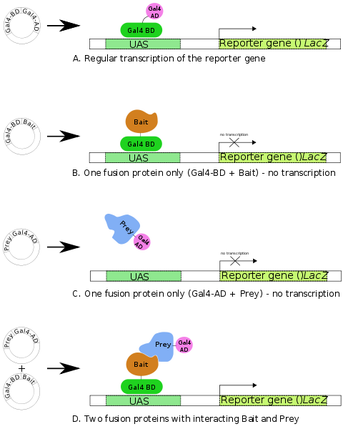
Figure 1. Diagram of YTH screen.
Protein-protein interactions occur in cells when one or more proteins bind together to carry out some kind of biological function. One common example of a biological function that occurs in all cells is DNA replication. This requires a large complex and involves many different types of proteins [1].
There are many ways to investigate these protein-protein interactions, two of which are yeast two hybrid analysis (YTH), and TAP tags. YTH screens can only be used in the nucleus, and are used to detect both interactions between two or more proteins, and protein and DNA. YTH screens work by fusing a bait protein to a DNA binding domain, and fusing a prey protein to an activation domain of a transcription factor [2]. This will then turn on a reporter gene of interest (see Figure 1). This shows that in eukaryotic cells, transcription factors are capable of activating transcription without directly binding to one another. This technique, when used in yeast cells, has helped researchers categorize many interacting proteins.
There are many ways to investigate these protein-protein interactions, two of which are yeast two hybrid analysis (YTH), and TAP tags. YTH screens can only be used in the nucleus, and are used to detect both interactions between two or more proteins, and protein and DNA. YTH screens work by fusing a bait protein to a DNA binding domain, and fusing a prey protein to an activation domain of a transcription factor [2]. This will then turn on a reporter gene of interest (see Figure 1). This shows that in eukaryotic cells, transcription factors are capable of activating transcription without directly binding to one another. This technique, when used in yeast cells, has helped researchers categorize many interacting proteins.
|
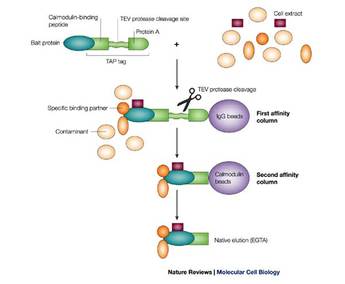
TAP tagging is the second method that will be discussed to investigate protein-protein interaction networks. TAP tags work by tagging the protein of interest with a TAP (Tandem Affinity Purification) tag so it binds with an IgG bead. The TAP tag is then cleaved at a specific site to allow the complex to bind to calmodulin coated beads [3]. The resulting complex is free of any contaminants. It has also been found that any bait protein in a specific complex will pull down the same proteins as any other protein in that complex, which means this technology is highly
|
reliable and reproducible [4]. TAP tags are also able to be used to screen for proteins outside of the nucleus, a limiting factor in YTH technology.
FRMD7 Protein-Protein Interactions
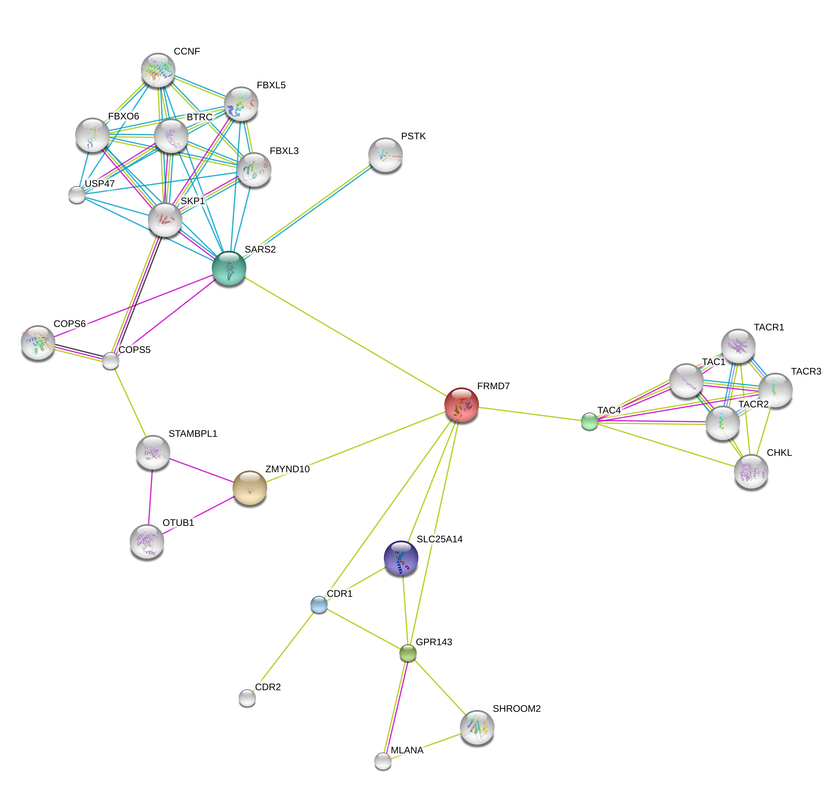
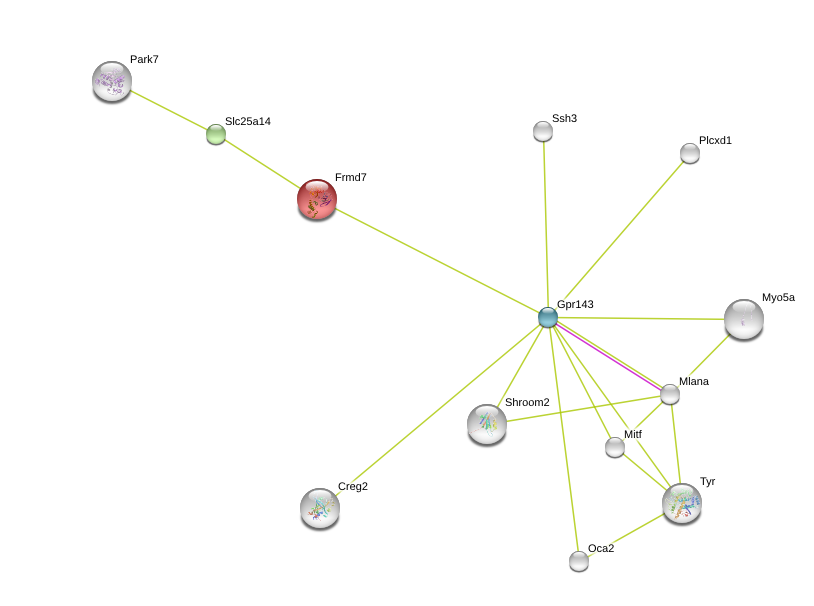
For this analysis of FRMD7, STRING was used to investigate its intracellular interaction. Both human and mice FRMD7 proteins were analyzed to explore differences between humans and model organisms. The interaction networks are below.
Analysis and Discussion
Functions of FRMD7 have yet to be well investigated by researchers to date, so this is probably the reason the interaction networks are so sparse. The first interesting thing to note is the interaction with TAC4, which is cleaved and secreted to excite neuronal cells. This may interact in some way with FRMD7 during growth cone formation in the embryo.
Another interesting thing that did come out of these networks is the connection mice FRMD7 has to Slc25a14 which in turn links to Park7, a gene involved in Parkinson's Disease. Slc25a14 has a homolog in humans, even though it does not directly interact with Park7, according to the above diagrams. In the future, it will be important to explore these protein interactions to determine if there are any connections between the diseases. This may aid in new drug treatments for nystagmus.
For more information on this topic, please visit the Conclusions and Future Directions page of this website.
Another interesting thing that did come out of these networks is the connection mice FRMD7 has to Slc25a14 which in turn links to Park7, a gene involved in Parkinson's Disease. Slc25a14 has a homolog in humans, even though it does not directly interact with Park7, according to the above diagrams. In the future, it will be important to explore these protein interactions to determine if there are any connections between the diseases. This may aid in new drug treatments for nystagmus.
For more information on this topic, please visit the Conclusions and Future Directions page of this website.
References
[1] Alberts, B., Johnson, A., Lewis J., Raff, M., Roberts, K., Walter, P. (2002)
Molecular Biology of the Cell, 4th Edition. New York: Garland Science, 2002. ISBN-10:0-8153-3218-1. Retrieved from: http://www.ncbi.nlm.nih.gov/books/NBK26856/#3215.
[2] Suter, B., Kittanakom, S., Stagljar, I. (2008)
Two-hybrid technologies in proteomics research. Current Opinion in Biotechnology. 2008 19(316). doi: 10.1016/j.copbio.1008.06.005.
[3] Collins, M. O., Choudhary, J. S. (2008)
Mapping multiprotein complexes by affinity purification and mass spectrometry. Current Opinion in Biotechnology. 2008 19(324). doi: 10.1016/j.copbio.2008.06.002.
[4] Gavin, AC., Bosche, M., Krause, R., Grandi, P., Marzioch, M., Bauer, A., Schultz, J., Rick, J. M., Michon A-M., Cruciat, C-M., Remor, M., Hofert, C., Schelder, M., Brajenovic, M., Ruffner, H., Merino, A., Klein, K., Hudak, M., Dickson, D., Rudi, T., Gnau, V., Gauch, A., Bastuck, S., Huhse, B., Leutwein, C., Heurtier, M-A., Copley, R., Edelmann, A., Querfurth, E., Rybin, V., Drewes, G., Raida, M., Bouwmeester, T., Bork, P., Seraphin, B., Kuster, B., Neubauer, G., Superti-Furga, G. (2002)
Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. 415(141). doi: 10.1038/415141a.
Molecular Biology of the Cell, 4th Edition. New York: Garland Science, 2002. ISBN-10:0-8153-3218-1. Retrieved from: http://www.ncbi.nlm.nih.gov/books/NBK26856/#3215.
[2] Suter, B., Kittanakom, S., Stagljar, I. (2008)
Two-hybrid technologies in proteomics research. Current Opinion in Biotechnology. 2008 19(316). doi: 10.1016/j.copbio.1008.06.005.
[3] Collins, M. O., Choudhary, J. S. (2008)
Mapping multiprotein complexes by affinity purification and mass spectrometry. Current Opinion in Biotechnology. 2008 19(324). doi: 10.1016/j.copbio.2008.06.002.
[4] Gavin, AC., Bosche, M., Krause, R., Grandi, P., Marzioch, M., Bauer, A., Schultz, J., Rick, J. M., Michon A-M., Cruciat, C-M., Remor, M., Hofert, C., Schelder, M., Brajenovic, M., Ruffner, H., Merino, A., Klein, K., Hudak, M., Dickson, D., Rudi, T., Gnau, V., Gauch, A., Bastuck, S., Huhse, B., Leutwein, C., Heurtier, M-A., Copley, R., Edelmann, A., Querfurth, E., Rybin, V., Drewes, G., Raida, M., Bouwmeester, T., Bork, P., Seraphin, B., Kuster, B., Neubauer, G., Superti-Furga, G. (2002)
Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature. 415(141). doi: 10.1038/415141a.
Site created by: Kristen Klimo
Last updated: 5/11/2012
University of Wisconsin-Madison
Last updated: 5/11/2012
University of Wisconsin-Madison