This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is Gene Ontology?
Gene Ontology (GO) is a major effort to unify the representation of a gene and gene product across all species where that gene or product (the protein) is present [1]. Currently in genetics, research is being done in thousands of laboratories, on many different model organisms, all over the world. Before the Gene Ontology project, every model organism had their own separate system for classifying genes and gene products, namely proteins. When labs would attempt to collaborate and look for homology among different species of model organisms, they found a surprising amount of similarity between the organisms. The first example of this is when the yeast and C. elegans genomes were compared, and researchers found that they had about 27% similarity between them [2]. They also found that transferring knowledge between one database to another was extremely difficult due to the different annotations each database used.
Today there are three categories of Gene Ontology. Molecular function is the first, and it is defined as the biological activity of a protein or gene product, as well as the capability that a protein or gene product could perform that biological activity [2]. A protein example of this is an enzyme. The second category is cellular component which is defined as the place in the cell where the protein or gene product is active. This is extremely important in eukaryotic cells because of subcellular compartmentalization. The third category is the biological process which refers to the biological objective to which a gene or protein contributes [2]. An example of this is proteins involved cellular growth and maintenance.
Today there are three categories of Gene Ontology. Molecular function is the first, and it is defined as the biological activity of a protein or gene product, as well as the capability that a protein or gene product could perform that biological activity [2]. A protein example of this is an enzyme. The second category is cellular component which is defined as the place in the cell where the protein or gene product is active. This is extremely important in eukaryotic cells because of subcellular compartmentalization. The third category is the biological process which refers to the biological objective to which a gene or protein contributes [2]. An example of this is proteins involved cellular growth and maintenance.
FRMD7 Protein Function
|
GO Terms
Biological Processes: Nervous system development Multicellular organismal developent Cellular Components: Cytoskeleton |
Molecular Functions: Growth cone development Neuron development |
|
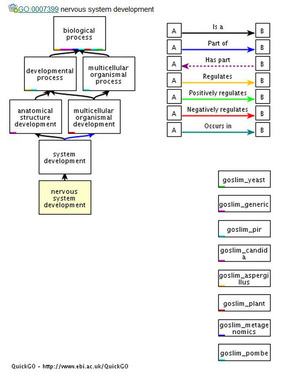
The flowchart to the right and the two below were obtained from QuickGo from the European Bioinformatics Institute. The flowcharts are meant to represent how the FRMD7 protein interacts in various pathways. FRMD7 is hypothesized to be involved with nervous system development, primarily in the region of the brain associated with ocular motor neurons [3]. The flowchart to the right is an overview for how the FRMD7 protein fits into the process of nervous system development.
|
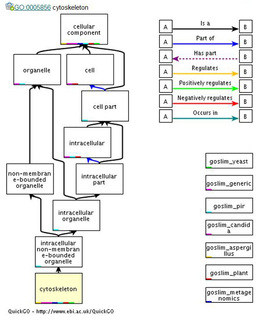
FRMD7 is also associated with components of the cytoskeleton, namely actin. In nerve cells, actin is extremely important for axonal growth cone development. Since it has been shown that FRMD7 co-localizes with actin, it is likely to also be involved with growth cone development [3]. Figure 2 below shows the how FRMD7 protein is involved with cytoskeleton development, and how this process fits into the neuron as a whole.
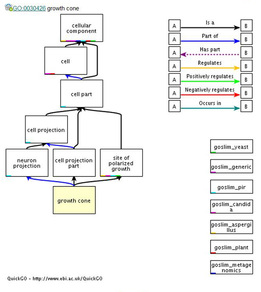
The growth cone is the extension of the neuron that will eventually become the axon, the end of the neuron that receives signals from the body or other neurons. Therefore, a defect in this process will cause the signals to be interrupted or disturbed, which is consistent with the disease phenotype presented by nystagmus patients. Figure 3 below shows the processes involved with growth cone development in the neuron cell.
For full results of of FRMD7 protein ontology, please click here.
The growth cone is the extension of the neuron that will eventually become the axon, the end of the neuron that receives signals from the body or other neurons. Therefore, a defect in this process will cause the signals to be interrupted or disturbed, which is consistent with the disease phenotype presented by nystagmus patients. Figure 3 below shows the processes involved with growth cone development in the neuron cell.
For full results of of FRMD7 protein ontology, please click here.
Analysis and Discussion
FRMD7 has yet to be well studied and understood. Since nystagmus is a defect in the motor neurons associated with eye movement, studies have been done to show FRMD7 protein is likely involved with growth cone development. If there is a problem with the growth cone, the signals the neuron cell should be receiving will be disrupted, which can cause the irregular eye movement of nystagmus.
References
[1] The Gene Ontology Project in 2008. Nucleic Acids Research, 2007(1). doi: 10.1093/nar/gkm883
[2] Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H., Cherry J. M., Davis A. P., Dolinski K., Dwight S. S., Eppig J. T., Harris M. A., Hill D. P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J. C., Richardson J. E., Ringwald M., Rubin G. M., Sherlock G. 2000
Gene Ontology: tool for the unification of biology. Nature Genetics, 25(1) doi: 10.1038/75556
[3] Betts-Henderson J., Bartesaghi, S., Crosier, M., Lindsay, S., Chen, H., Salomoni, P., Gottlob, I., Nicotera P. (2010)
The nystagmus-associated FRMD7 gene regulates neuronal outgrowth and development. Human Molecular Genetics, 19(2), 342. doi: 10.1093/hmg/ddp500
[2] Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H., Cherry J. M., Davis A. P., Dolinski K., Dwight S. S., Eppig J. T., Harris M. A., Hill D. P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J. C., Richardson J. E., Ringwald M., Rubin G. M., Sherlock G. 2000
Gene Ontology: tool for the unification of biology. Nature Genetics, 25(1) doi: 10.1038/75556
[3] Betts-Henderson J., Bartesaghi, S., Crosier, M., Lindsay, S., Chen, H., Salomoni, P., Gottlob, I., Nicotera P. (2010)
The nystagmus-associated FRMD7 gene regulates neuronal outgrowth and development. Human Molecular Genetics, 19(2), 342. doi: 10.1093/hmg/ddp500
Site created by: Kristen Klimo
Last updated: 5/11/2012
University of Wisconsin-Madison
Last updated: 5/11/2012
University of Wisconsin-Madison