This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Motifs and Domains
One of the most common misconceptions about the words "domain" and "motif" is that they are interchangeable. For genes, the word domain refers to a piece of the overall gene that gives rise to a functional unit capable of functioning on it's own without the entire structure. A gene motif is the actual nucleotide sequence that results in a functional unit. These domains are often similar or conserved across many different taxa.
There are three main types of DNA sequence motif discovery, enumeration, deterministic optimization, and probabilistic optimization [1]. Enumeration uses complex algorithms to cover all possible motifs looking for a specific motif model description. This technique counts the number of occurrences of a particular sequence to calculate which ones are overrepresented. This technique is often used to find consensus sequences for a particular site. These consensus sequences allow for a number of mismatches to pinpoint target sequences.
Deterministic optimization is the technique used by MEME, and performs a single iteration for each n-mer in the target sequences. It then selects the best motif from this set and repeats only that one to convergence. This provides some assurance that the algorithm is unlikely to get stuck at a poor local maximum [1].
The third type of DNA sequence motif is probabilistic optimization, or Gibbs sampling. This method uses a randomly selected set of sites with target sequences that are scored against an initial motif model. The algorithm then uses probability to decide whether to add or remove an old site from this model using binding probability for those sites [1].
The majority of gene product proteins are are multidomain proteins meaning they have more than one functional domain present in their structure [2]. These multiple domains probably existed independently at some point during the evolutionary timeline. Multidomain proteins arise from transposition of mobile genetic elements, large scale gene mutations - deletions, duplication, translocations etc. - homologous recombination and DNA polymerase replication errors [3].
There are three main types of DNA sequence motif discovery, enumeration, deterministic optimization, and probabilistic optimization [1]. Enumeration uses complex algorithms to cover all possible motifs looking for a specific motif model description. This technique counts the number of occurrences of a particular sequence to calculate which ones are overrepresented. This technique is often used to find consensus sequences for a particular site. These consensus sequences allow for a number of mismatches to pinpoint target sequences.
Deterministic optimization is the technique used by MEME, and performs a single iteration for each n-mer in the target sequences. It then selects the best motif from this set and repeats only that one to convergence. This provides some assurance that the algorithm is unlikely to get stuck at a poor local maximum [1].
The third type of DNA sequence motif is probabilistic optimization, or Gibbs sampling. This method uses a randomly selected set of sites with target sequences that are scored against an initial motif model. The algorithm then uses probability to decide whether to add or remove an old site from this model using binding probability for those sites [1].
The majority of gene product proteins are are multidomain proteins meaning they have more than one functional domain present in their structure [2]. These multiple domains probably existed independently at some point during the evolutionary timeline. Multidomain proteins arise from transposition of mobile genetic elements, large scale gene mutations - deletions, duplication, translocations etc. - homologous recombination and DNA polymerase replication errors [3].
Motifs in FRMD7
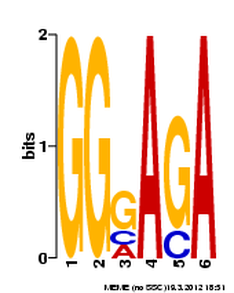
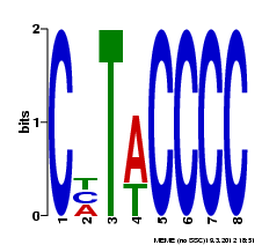
For this analysis, MEME was used to compare human, dog, rat, chicken and cattle FRMD7 gene fragments. The entire gene sequence was too long for the program, so only the portion encoding FERM domain was used. This particular portion of the gene was used because it is the most common region with mutations associated with nystagmus. The results from the MEME analysis are below.
Analysis and Discussion
The results above, obtained from MEME, show three different DNA motifs present in FRMD7. These results are fairly easy to interpret, where the size of the nucleotide shows the prevalence in the DNA. The larger the nucleotide, the more often it is present when comparing sequences. MEME allows for some differences, which result in the double listing of nucleotides.
The first two of these sequences are very short, which doesn't tell researchers much about their function or importance. The third one, while it has quite a bit of variation among species, is much longer with a e-value of 6.8e+002.
It would have probably been more helpful to be able to enter the entire FRMD7 sequence for more useful results. This also would have accounted for changes that occur outside of the FERM domain part of the gene.
The first two of these sequences are very short, which doesn't tell researchers much about their function or importance. The third one, while it has quite a bit of variation among species, is much longer with a e-value of 6.8e+002.
It would have probably been more helpful to be able to enter the entire FRMD7 sequence for more useful results. This also would have accounted for changes that occur outside of the FERM domain part of the gene.
References
[1] D'haeseleer, P. (2006)
How does DNA sequence motif discovery work? Nature Biotechnology. 24(8), 959. doi: 10.1038/nbt0806-959
[2] Apic, G., Gough, J., and Teichmann, S. A. (2001).
Domain combinations in archaeal, eubacterial and eukaryotic proteomes. Journal of Molecular Biology, 310(2), 311. doi: 10.1006/jmbi.2001.4776
[3] Bork, P. and Doolittle, R. F (1992).
Proposed aquisition of animal protein domain by bacteria. Proceeding of the National Adademy of Science. 89(19), 8990. doi: 10.1073/pnas.8919.8990
How does DNA sequence motif discovery work? Nature Biotechnology. 24(8), 959. doi: 10.1038/nbt0806-959
[2] Apic, G., Gough, J., and Teichmann, S. A. (2001).
Domain combinations in archaeal, eubacterial and eukaryotic proteomes. Journal of Molecular Biology, 310(2), 311. doi: 10.1006/jmbi.2001.4776
[3] Bork, P. and Doolittle, R. F (1992).
Proposed aquisition of animal protein domain by bacteria. Proceeding of the National Adademy of Science. 89(19), 8990. doi: 10.1073/pnas.8919.8990
Site created by: Kristen Klimo
Last updated: 5/11/2012
University of Wisconsin-Madison
Last updated: 5/11/2012
University of Wisconsin-Madison